Revealing hidden genomic complexity in AAV manufacturing using the NanoMosaic platform
Uncovering hidden heterogeneity in rAAV genomes
Recombinant AAV (rAAV) vector preparations are rarely uniform. Beneath the surface lies a mixture of full-length, partially packaged, and truncated genomes. Each of these species can influence potency, safety, and yield in distinct ways. Accurate quantification of these heterogeneous species is critical for understanding and improving product quality, as well as meeting regulatory expectations for genome integrity.
Primer-based strategies to map genome integrity
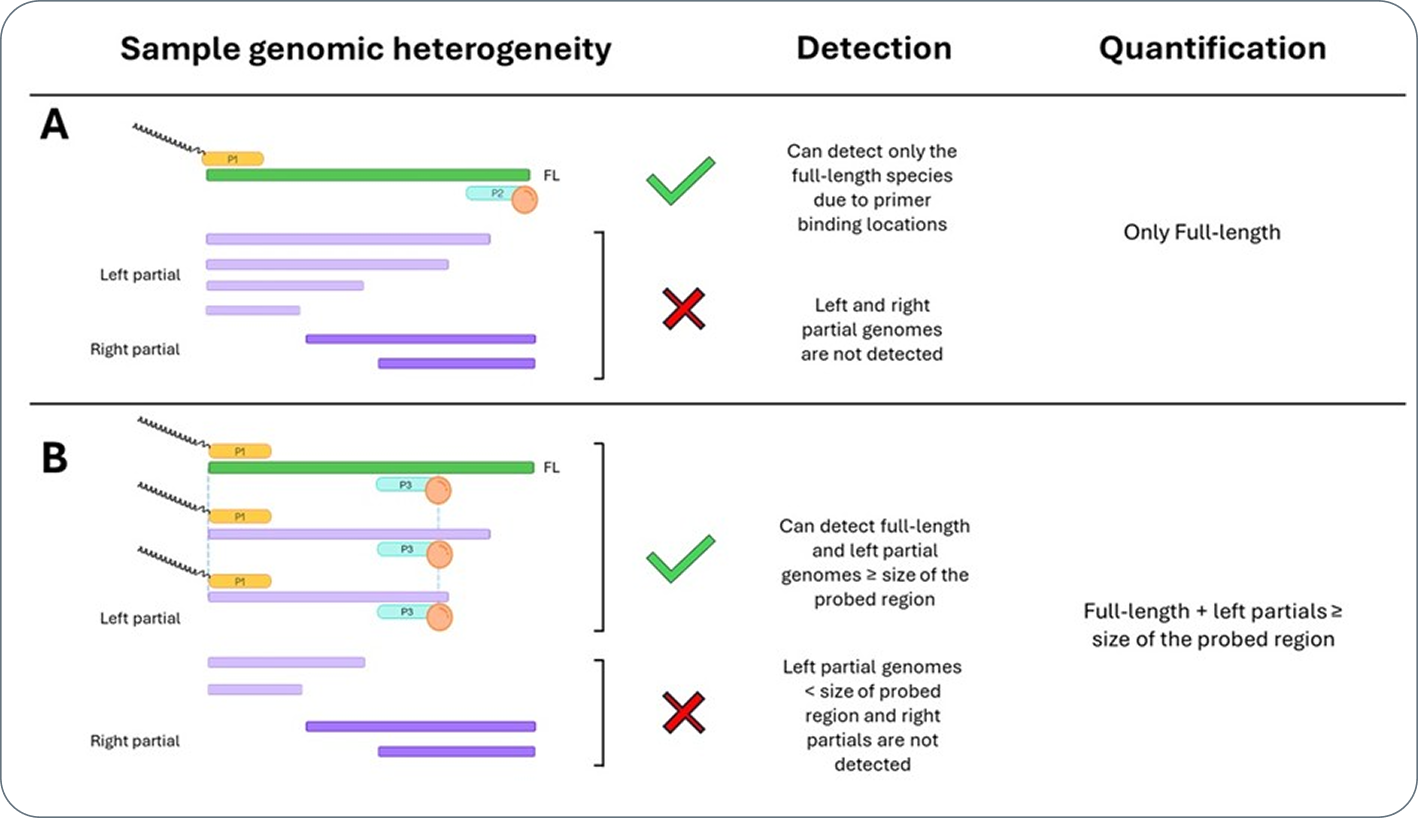
The figure below illustrates how primer-based assays can help untangle this complexity.
In the full-length assay (Panel A), primers positioned at the 5′ and 3′ ends (P1 and P2) amplify only intact genomes, effectively counting the therapeutically relevant, complete species.
To better assess transgene regions contributing to truncations or partial genomes, the Probe Walk strategy can be applied. In Panel B, the forward primer (P1) remains in the same position as in the full-length assay, while the reverse primer (P3) is shifted inward. This configuration captures both full-length and left-partial genomes that extend beyond the probed region, enabling selective detection of truncated variants and mapping of truncation hotspots.
From detection to optimization: improving vector design & production
This nuanced detection framework, in which amplification occurs only when both primer sites are present, offers more than a means of quality control. It provides a diagnostic window into process biology. By pinpointing where truncations occur, the Probe Walk assay informs rational sequence engineering (for example, removing secondary structures or GC-rich regions prone to breakage) and supports upstream process optimization to mitigate genome fragmentation during replication or packaging.
We applied this strategy to an rAAV9 vector carrying a CAG-Luciferase-WPRE-bGH p(A) transgene purified by cesium chloride gradient. The analysis revealed distinct truncation patterns, highlighting actionable regions for vector redesign and improved manufacturing consistency.
To read more about that, see our preprint in bioRxiv: https://doi.org/10.64898/2026.03.03.709319
Revealing hidden genomic complexity in AAV manufacturing using the NanoMosaic platform

Uncovering hidden heterogeneity in rAAV genomes
Recombinant AAV (rAAV) vector preparations are rarely uniform. Beneath the surface lies a mixture of full-length, partially packaged, and truncated genomes. Each of these species can influence potency, safety, and yield in distinct ways. Accurate quantification of these heterogeneous species is critical for understanding and improving product quality, as well as meeting regulatory expectations for genome integrity.
Primer-based strategies to map genome integrity
The figure below illustrates how primer-based assays can help untangle this complexity.
In the full-length assay (Panel A), primers positioned at the 5′ and 3′ ends (P1 and P2) amplify only intact genomes, effectively counting the therapeutically relevant, complete species.
To better assess transgene regions contributing to truncations or partial genomes, the Probe Walk strategy can be applied. In Panel B, the forward primer (P1) remains in the same position as in the full-length assay, while the reverse primer (P3) is shifted inward. This configuration captures both full-length and left-partial genomes that extend beyond the probed region, enabling selective detection of truncated variants and mapping of truncation hotspots.

From detection to optimization: improving vector design & production
This nuanced detection framework, in which amplification occurs only when both primer sites are present, offers more than a means of quality control. It provides a diagnostic window into process biology. By pinpointing where truncations occur, the Probe Walk assay informs rational sequence engineering (for example, removing secondary structures or GC-rich regions prone to breakage) and supports upstream process optimization to mitigate genome fragmentation during replication or packaging.
We applied this strategy to an rAAV9 vector carrying a CAG-Luciferase-WPRE-bGH p(A) transgene purified by cesium chloride gradient. The analysis revealed distinct truncation patterns, highlighting actionable regions for vector redesign and improved manufacturing consistency.
To read more about that, see our preprint in bioRxiv: https://doi.org/10.64898/2026.03.03.709319
