Resolving The Hidden Gap Between AAV Genome Titer and Potency
The NanoMosaic platform enables measurement of therapeutically relevant viral genome titer in AAV QC analytics and is the first and only FDA/CBER-designated Advanced Manufacturing Technology for this purpose.
AAV manufacturing, vector genome (VG) titer is often treated as a proxy for dose. However, not all genome copies are equal. Conventional short-amplicon qPCR assays quantify a small internal region of the transgene, implicitly assuming that detection of that region reflects the presence of a complete, therapeutically functional genome.
The figure below highlights a critical discrepancy. When a short internal region is measured, the apparent titer can be approximately 20-fold higher than the number of full-length viral genomes capable of expressing the intended transgene.
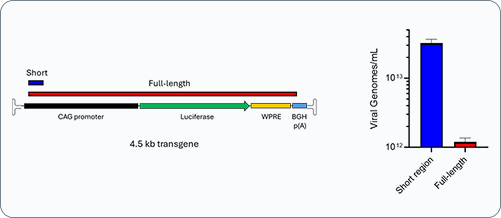
Why does this happen?
Short-region assays detect any genome fragment that contains the probed sequence. This includes partially packaged, truncated, or fragmented species that retain the internal region but lack essential 5′ or 3′ elements required for functional expression. As a result, short-region titer can substantially overestimate the number of therapeutically competent vectors in the preparation.
In the context of AAV gene therapy manufacturing, this gap has meaningful implications:
- Potency misestimation: Dosing decisions may be based on inflated VG count
- Process optimization blind spots: Truncation patterns and packaging inefficiencies remain invisible.
- Regulatory scrutiny: Genome integrity is increasingly expected to be characterized beyond a single internal amplicon.
The unmet need is clear: we must distinguish total genome fragments from true full-length, expression-competent vectors.
NanoMosaic assays address this directly by enabling full-length genome quantification and truncation mapping
in the same analytical framework. By strategically positioning primers across the transgene, our approach differentiates intact genomes from partial species and reveals where fragmentation occurs. This transforms titer from a single number into a biologically meaningful quality attribute.
By resolving this distinction, manufacturers gain actionable insight for sequence redesign, capsid packaging optimization, and upstream process refinement - closing the gap between apparent titer and true therapeutic potential.
We applied this strategy to an rAAV9 vector carrying a CAG-Luciferase-WPRE-bGH p(A) transgene purified by cesium chloride gradient.
To read more about that, find the full publication on bioRxiv.
Resolving The Hidden Gap Between AAV Genome Titer and Potency
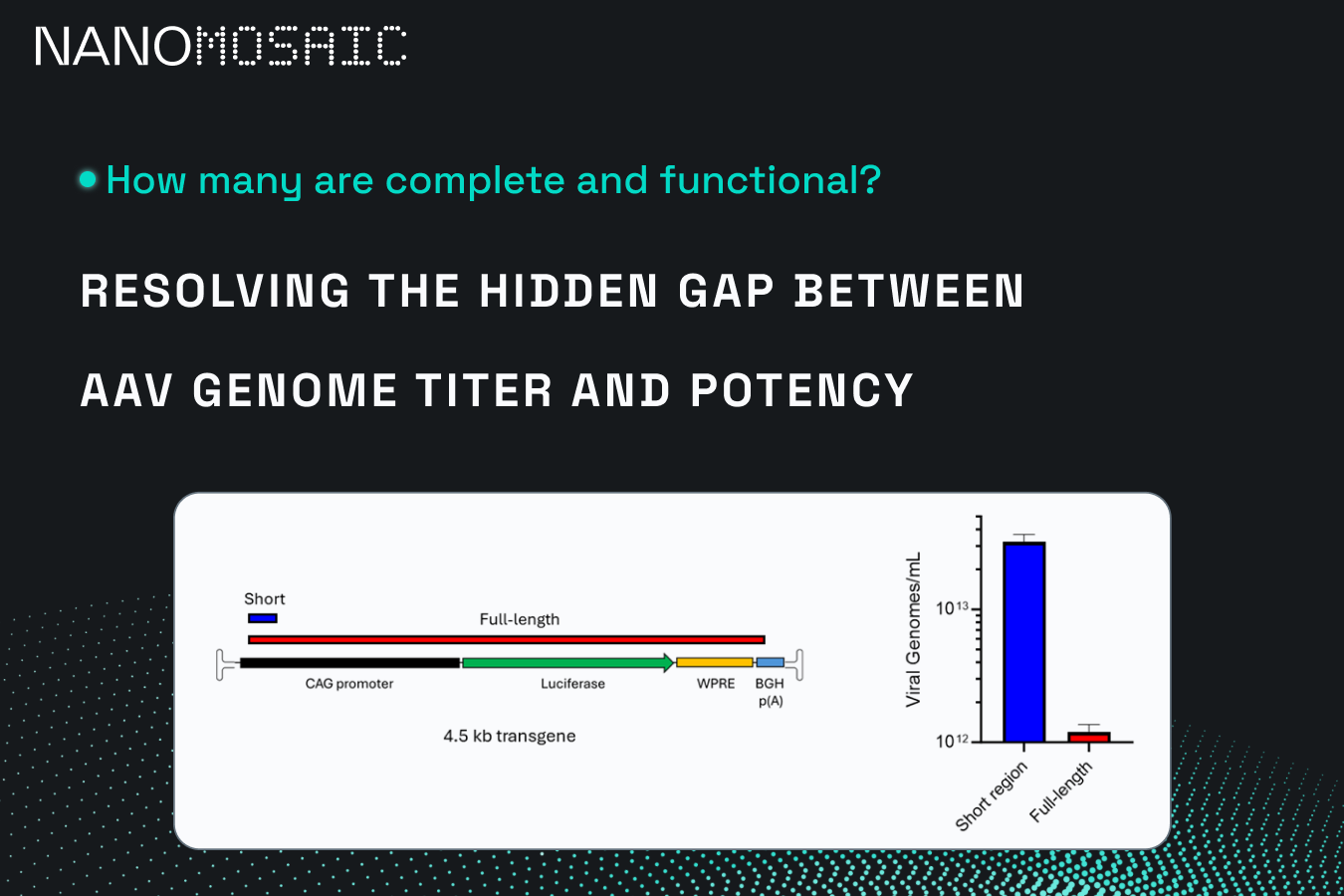
The NanoMosaic platform enables measurement of therapeutically relevant viral genome titer in AAV QC analytics and is the first and only FDA/CBER-designated Advanced Manufacturing Technology for this purpose.
AAV manufacturing, vector genome (VG) titer is often treated as a proxy for dose. However, not all genome copies are equal. Conventional short-amplicon qPCR assays quantify a small internal region of the transgene, implicitly assuming that detection of that region reflects the presence of a complete, therapeutically functional genome.
The figure below highlights a critical discrepancy. When a short internal region is measured, the apparent titer can be approximately 20-fold higher than the number of full-length viral genomes capable of expressing the intended transgene.

Why does this happen?
Short-region assays detect any genome fragment that contains the probed sequence. This includes partially packaged, truncated, or fragmented species that retain the internal region but lack essential 5′ or 3′ elements required for functional expression. As a result, short-region titer can substantially overestimate the number of therapeutically competent vectors in the preparation.
In the context of AAV gene therapy manufacturing, this gap has meaningful implications:
- Potency misestimation: Dosing decisions may be based on inflated VG count
- Process optimization blind spots: Truncation patterns and packaging inefficiencies remain invisible.
- Regulatory scrutiny: Genome integrity is increasingly expected to be characterized beyond a single internal amplicon.
The unmet need is clear: we must distinguish total genome fragments from true full-length, expression-competent vectors.
NanoMosaic assays address this directly by enabling full-length genome quantification and truncation mapping
in the same analytical framework. By strategically positioning primers across the transgene, our approach differentiates intact genomes from partial species and reveals where fragmentation occurs. This transforms titer from a single number into a biologically meaningful quality attribute.
By resolving this distinction, manufacturers gain actionable insight for sequence redesign, capsid packaging optimization, and upstream process refinement - closing the gap between apparent titer and true therapeutic potential.
We applied this strategy to an rAAV9 vector carrying a CAG-Luciferase-WPRE-bGH p(A) transgene purified by cesium chloride gradient.
To read more about that, find the full publication on bioRxiv.
